This project will explore the possibilities for implementing creative evolution in artificial life software, responding to the challenge by Bedau et al (2000) to reproduce "open-ended evolution". We will begin by examining previous attempts and Bedau's evolutionary activity statistics (Bedau et al 1998) for assessing them. We will develop our own measures of the growth of complexity in
organism structure and behaviour, and apply them to:
- The paleontological record,
- The results of existing Artificial Life simulations, and
- New simulations based on existing work by Yaeger (2006) and the supervisors.
Students are requested to do some preliminary reading (e.g. at least (Levy 1997) and (Bedau et al 2000)) prior to commencing the project. Enrollment in the Honours unit FIT4012 is strongly encouraged as it will provide background material of direct relevance to the topic.
References:
- Bedau, M.A., et al., Open Problems in Artificial Life. Artificial Life, 2000. 6(4): p. 363-376.
- Bedau, M.A., E. Snyder, and N.H. Packard. A Classification of Long-Term Evolutionary Dynamics. In Artificial Life VI. 1998: MIT Press: p. 228-237.
- Levy, S., Artificial Life, the quest for a new creation. 1992, London: Penguin Books. 390
- Yaeger, L.S., O. Sporns. Evolution of Neural Structure and Complexity in a Computational Ecology. In Artificial Life X. Rocha, L. et al. (eds.) Cambridge, MA: MIT Press. 2006.

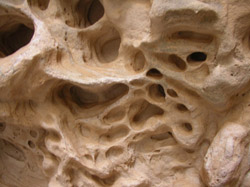
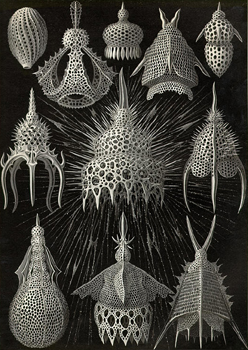
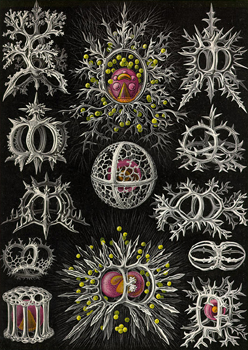
The components of many organisms and their secretions consist of what may loosely be labelled "sinewy networks". The junctions in these networks are often gently filleted, creating a web-like mesh of holes and curves that gives the structure a biological feel. The purpose of this project is to devise a technique for automating the process of generating the geometry of these meshes. The software will be parameterized to vary the extent to which the meshes are filletted, allowing them to resemble architectural forms of different kinds, whether they be made by spiders (webs), erosion (rocks hollows and caves), plants (patterns in bark, veins on leaves, tree roots) or people (architectural lattices). The software will also allow automatic generation of the network itself and should allow the user to parameterize the form of the lattice in general terms from completely irregular, through approximately regular, to highly geometric.
Some visual examples of the kinds of structure the project aims to build are given below. (Photographs © Copyright Alan Dorin 2005)