|
This
document is online at
users.monash.edu/~lloyd/Seminars/2001-WEHI/index.shtml
and contains hyper-links to more detailed notes on certain topics
and to some of the references. |
|
The story...
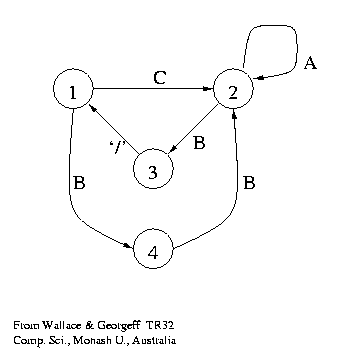
A very (!) old problem:
Learn a language.
Model Complexity:
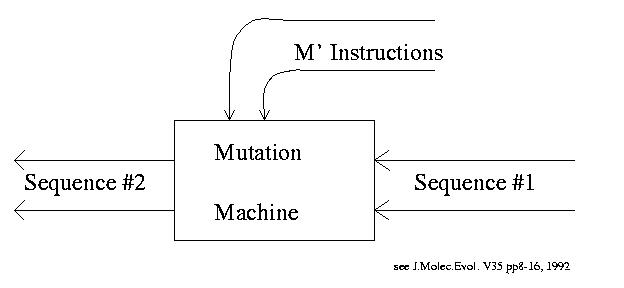
Mutation Machine

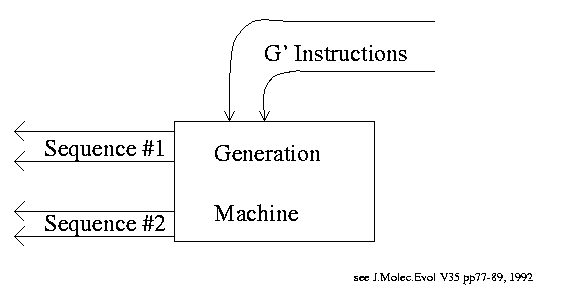
Equivalent Generation Machine:
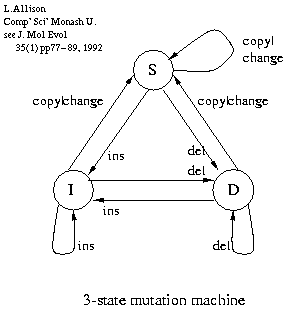
Compression ~ Sequence Complexity:
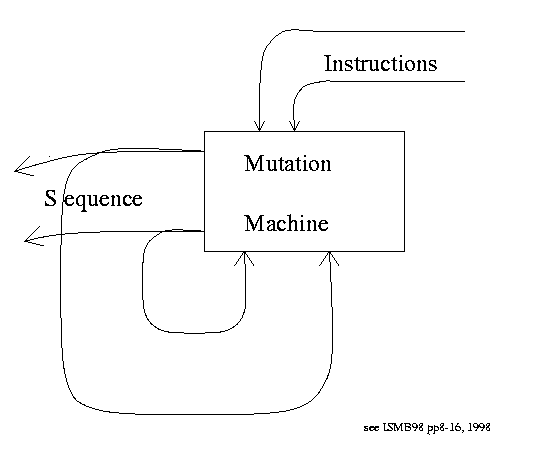
What matters most is the control box, e.g.
 |
p.falciparum, chr2, 0.95Mbp [colour] [b+w] [1-D plot] bits/b.p. NB. Same twist - sum over all explanations. But not enough space for fwd+rev for long Seq'. |
| Seq_1 | . . . | S1[i] | . . . | |
|---|---|---|---|---|
| S e q 2 . |
||||
| . . . |
.
(mis)match .
.
.
|
| |
||
| S2[j] | |
------->
| ? | |
| . . . |